Determination of low strain interfaces via geometric matching¶
Version: 2017.1
This tutorial introduces the new GeneralizedLatticeMatch method for combining two bulk crystals into an interface.
On many occasions, one is interested in creating a realistic interface between two materials, even without having precise structural informations on the two surfaces that form the interface. The GeneralizedLatticeMatch method facilitates this process, by automatically finding all the possible interface supercells between the two crystals based only on their bulk crystalline structure. The method is an optimized version of the algortihm described in [1]. Compared to the lattice-matching method used in the Interface Builder, described in Interface Builder, the present method is unbiased as it considers all the possible surfaces of the two crystals forming the interface.
A number of structural parameters are calculated for each interface, which allows one to easily analyze the matching pattern and choose the most appropriate interface to be used for further studies. The structure of the desired interface can then be easily created using the Interface Builder implemented in QuantumATK.

Method description¶
The following describes in the main steps of the algortihm used in the GeneralizedLatticeMethod method, which is very similar to that used in the Interface Builder (see Interface Builder).
Initially, the possible surface vectors \([\mathbf{a}_{1}, \mathbf{a}_{2}]\) of the first of the two materials forming the interface are constructed, starting from a linear combination of the Bravais vectors of the primitive cell:
\[ \begin{align}\begin{aligned}\mathbf{a}_1 = \sum_{i=1}^3 c_i \mathbf{u}_i \quad c_i \in \mathbb{Z},\\\mathbf{a}_2 = \sum_{i=1}^3 c_i^\prime \mathbf{u}_i \quad c_i^\prime \in \mathbb{Z},\end{aligned}\end{align} \]and by a subsequent projection of the resulting vectors from \(\mathbf{R}^3\) to \(\mathbf{R}^2\). A similar procedure is applied to construct the surface vectors \([\mathbf{b}_{1}, \mathbf{b}_{2}]\) of the second surface. As it will be shown in the Section Input and output description, the number of generated surface cells can be limited by specifying:
The maximum value of the Miller indexes \([h,k,l]\) that define each surface;
The maximum length of the lattice vectors \([\mathbf{a}_{1}, \mathbf{a}_{2}]\) and \([\mathbf{b}_{1}, \mathbf{b}_{2}]\).
Each couple of surface cells is matched by using the same lattice-match method as used in the Interface Builder, and described in the technical note Interface Builder.
The average strain is then calculated as:
\[\mathbf{\bar{\varepsilon}} = \sqrt{\frac{\varepsilon_{11}^2 + \varepsilon_{22}^2 + \varepsilon_{11}\varepsilon_{22} + \varepsilon_{12}^2}{4}}\]where \(\epsilon_{11}\), \(\epsilon_{22}\) and \(\epsilon_{12}\) are the components of the 2D strain tensor, as defined in the technical note Interface Builder.
Note
Notice that this definition of the average strain differs from that used in the Interface Builder. The present definition of strain is more appropriate for the present method as it is an invariant of the strain tensor.
Input and output description¶
In order to use the GeneralizedLatticeMatch method one can set up a simple script as follows:
1# Read the BulkConfiguration of the primitive cell of the
2# first material
3configuration_1 = nlread('InAs.py',BulkConfiguration)[-1]
4# Read the BulkConfiguration of the primitive cell of the
5# second material
6configuration_2 = nlread('Al.py',BulkConfiguration)[-1]
7
8# Run the GeneralizedLatticeMatch method
9generalized_lattice_match = GeneralizedLatticeMatch(
10 configuration_1,
11 configuration_2,
12 max_strain=0.02,
13 maximum_miller_index=3,
14 longest_surface_lattice_vector=50*Angstrom,
15 max_surface_area=200.0*Angstrom**2,
16 user_given_miller_index=None
17 )
The script reads two input files, each one containing a BulkConfiguration of the bulk primitive cell of one of the two materials that form the interface. Then, it applies the GeneralizedLatticeMatch method on these two structures. A number of input parameters can be set to control the precision and the extent of the search for possible interface supercells. The full list of input parameters is:
configuration_1: BulkConfiguration of the bulk primitive cell of the first material.configuration_2: BulkConfiguration of the bulk primitive cell of the second material.max_strain: Maximum strain applied to each of the two surfaces.maximum_miller_index: Maximum value of the Miller indexes \(\left[ h, k, l \right]\) that define each of the two surfaces.longest_surface_lattice_vector: Maximum length of each surface lattice vector \([\mathbf{a}_{1}, \mathbf{a}_{2}]\) and \([\mathbf{b}_{1}, \mathbf{b}_{2}]\).max_surface_area: Maximum value of the surface area of the interface supercell.user_given_miller_index: Predefined Miller indexes of the surface ofconfiguration_1.Note
When the parameter
user_given_miller_indexis set to a value different thanNone, the search for the possible surfaces is restricted to the second material, whereas the surface of the first material is kept fixed. This option will be used in Example 2: Lattice match between a bulk system with a predefined surface.
After the possible matches are calculated, the matching results are printed directly in the QuantumATK log file of the calculation. The matches will be listed in order of increasing strain, together with a number of output parameters that characterize each match.
+----------------------------------------------------------------------------------------+
| A B Strain Atoms Area Aspect Angle Rotation |
+----------------------------------------------------------------------------------------+
[ 1 0 0] >-< [ 1 0 0] 0.000110 29 72.9 1.0 90.0 0.0
[ 2 2 1] >-< [ 1 0 0] 0.000110 23 163.9 2.2 153.4 26.6
[ 3 3 2] >-< [ 1 1 1] 0.004030 28 170.9 1.2 162.6 60.3
The output parameters are:
A: Miller indexes \(\left[h, k, l \right]\) of the first surface.
B: Miller indexes \(\left[h, k, l \right]\) of the second surface.
Strain: Maximum strain of the two surfaces.
Atoms: Total number of atoms in the interface supercell.
Area: Surface area of the interface supercell.
Aspect: Aspect ratio between two surface vectors of the interface supercell. The ratio is calculated by taking the largest vector at the numerator.
Angle: Angle between the two vectors of the interface supercell.
Rotation: Rotation between the two surfaces in the interface supercell.
Example 1: Lattice match between two bulk systems¶
In this first example, you will calculate the possible matches between indium arsenide and aluminum.
To obtain the structure file containing the BulkConfiguration of bulk InAs,
open QuantumATK, go to the  Builder and click on
. In the Database, select the
primitive cell of bulk InAs shown in the figure below, add it to the Stash
by clicking on the
Builder and click on
. In the Database, select the
primitive cell of bulk InAs shown in the figure below, add it to the Stash
by clicking on the  button.
button.
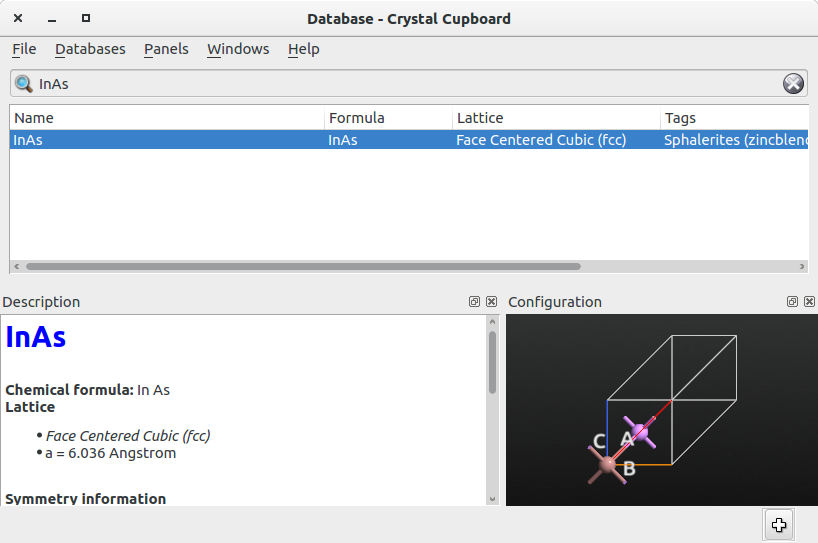
Repeat the same procedure to add to the Stash the structure of the primitive cell of bulk Al, which is shown in the figure below.
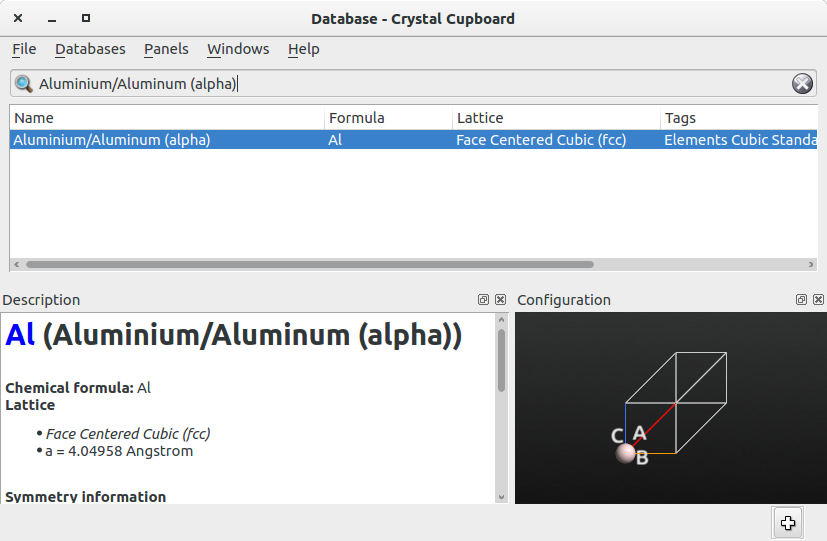
In the Stash, save the two configurations by right-clicking
on each structure and selecting ‘Save as’. Alternatively, you can download
the two files from here: InAs.py, Al.py.
Set up the script for the GeneralizedLatticeMatch method as shown below:
1# Read the BulkConfiguration of the primitive cell of the
2# first material
3configuration_1 = nlread('InAs.py',BulkConfiguration)[-1]
4# Read the BulkConfiguration of the primitive cell of the
5# second material
6configuration_2 = nlread('Al.py',BulkConfiguration)[-1]
7
8# Run the GeneralizedLatticeMatch method
9generalized_lattice_match = GeneralizedLatticeMatch(
10 configuration_1,
11 configuration_2,
12 max_strain=0.02,
13 maximum_miller_index=3,
14 longest_surface_lattice_vector=50*Angstrom,
15 max_surface_area=200.0*Angstrom**2,
16 user_given_miller_index=None
17 )
Alternatively, you can download the script from here: match_InAs_Al.py
In this example, the scan will be performed by considering surfaces with a maximum strain of 0.02, with Miller indexes \(h,k,l \leq 3\), with a maximum length of the lattice vectors of 50 Å, and with a upper threshold for the surface area of 200 Å2.
Run the script in the terminal as atkpython match_InAs_Al.py > match_InAs_Al.log.
The output will look as shown below. There are several possible matches, so
for briefness only the first 20 matches are shown.
+------------------------------------------------------------------------------+
| |
| Atomistix ToolKit 2017.1 [Build ce08f12] |
| |
+------------------------------------------------------------------------------+
|--------------------------------------------------|
Miller planes for A : ==================================================
|--------------------------------------------------|
Miller planes for B : ==================================================
|--------------------------------------------------|
Matching Miller planes : ==================================================
+----------------------------------------------------------------------------------------+
| A B Strain Atoms Area Aspect Angle Rotation |
+----------------------------------------------------------------------------------------+
[ 1 0 0] >-< [ 1 0 0] 0.000110 29 72.9 1.0 90.0 0.0
[ 2 2 1] >-< [ 1 0 0] 0.000110 23 163.9 2.2 153.4 26.6
[ 3 3 2] >-< [ 1 1 1] 0.004030 28 170.9 1.2 162.6 60.3
[ 3 1 1] >-< [ 2 1 1] 0.004030 10 120.8 1.7 90.0 0.0
[ 3 1 1] >-< [ 3 1 1] 0.004250 16 120.8 1.7 106.8 33.6
[ 3 3 1] >-< [ 3 3 1] 0.004250 16 158.8 2.2 102.9 0.0
[ 3 3 1] >-< [ 3 3 1] 0.005470 13 158.8 2.2 102.9 0.0
[ 3 1 1] >-< [ 3 1 1] 0.005470 13 120.8 1.7 106.8 33.6
[ 2 1 1] >-< [ 2 1 1] 0.005470 17 178.5 2.4 90.0 0.0
[ 2 1 0] >-< [ 2 1 0] 0.005470 13 162.9 1.2 114.1 0.0
[ 1 1 1] >-< [ 1 1 1] 0.005470 13 63.1 1.0 120.0 60.0
[ 1 1 0] >-< [ 1 1 0] 0.005470 17 103.0 1.4 90.0 0.0
[ 1 0 0] >-< [ 1 0 0] 0.005470 13 72.9 1.0 90.0 0.0
[ 1 0 0] >-< [ 2 2 1] 0.005470 7 72.9 1.0 90.0 0.0
[ 3 2 2] >-< [ 1 1 1] 0.005940 23 150.2 1.2 157.6 60.5
[ 1 0 0] >-< [ 2 1 1] 0.006130 19 182.2 3.9 39.8 0.6
[ 3 1 1] >-< [ 1 1 0] 0.007160 18 151.0 1.7 73.2 22.6
[ 3 1 1] >-< [ 1 0 0] 0.007730 19 120.8 1.7 90.0 0.0
[ 3 2 2] >-< [ 3 1 1] 0.008250 13 150.2 2.3 115.9 35.2
[ 1 0 0] >-< [ 1 1 0] 0.008420 28 200.4 2.2 79.7 38.2
Notice how the match with the least strain is that between the \(\langle 100 \rangle\) surface of InAs and the \(\langle 100 \rangle\) surface of Al.
Example 2: Lattice match between a bulk system with a predefined surface¶
In some cases, one could be interested in finding the possible matches of a bulk material to a given surface. In this second example, you will investigate how to find the possible matches of bulk aluminium to the InAs(111) surface. InAs nanowires with \(\langle 111 \rangle\)-oriented facets have been synthesized in Ref. [2], and it has been demonstrated experimentally that \(\langle 111 \rangle\)-oriented Al layers can be grown on this surface.
Use the same structure files as in Example 1: Lattice match between two bulk systems, and setup the script for the GeneralizedLatticeMatch calculation as follows:
1# Read the BulkConfiguration of the primitive cell of the
2# first material
3configuration_1 = nlread('InAs.py',BulkConfiguration)[-1]
4# Read the BulkConfiguration of the primitive cell of the
5# second material
6configuration_2 = nlread('Al.py',BulkConfiguration)[-1]
7
8# Run the GeneralizedLatticeMatch method
9generalized_lattice_match = GeneralizedLatticeMatch(
10 configuration_1,
11 configuration_2,
12 max_strain=0.02,
13 maximum_miller_index=3,
14 longest_surface_lattice_vector=50*Angstrom,
15 max_surface_area=200.0*Angstrom**2,
16 user_given_miller_index=(1,1,1)
17 )
Notice how the user_given_miller_index option, highlighted in yellow, has now
been set to user_given_miller_index=(1,1,1). Therefore, in this case only the
matches of bulk Al to the InAs(111) surface will be considered. You can also download
the script from here: match_InAs111_Al.py
Run the script in the terminal as atkpython match_InAs111_Al.py > match_InAs111_Al.log.
The output will look as shown below. Notice how there are fewer possible matches
than those in the output of Example 1: Lattice match between two bulk systems. This is because only one surface is
now considered for the material denoted A.
+------------------------------------------------------------------------------+
| |
| Atomistix ToolKit 2017.1 [Build ce08f12] |
| |
+------------------------------------------------------------------------------+
|--------------------------------------------------|
Miller planes for B : ==================================================
|--------------------------------------------------|
Matching Miller planes : ==================================================
+----------------------------------------------------------------------------------------+
| A B Strain Atoms Area Aspect Angle Rotation |
+----------------------------------------------------------------------------------------+
[ 1 1 1] >-< [ 1 1 1] 0.005470 13 63.1 1.0 120.0 60.0
[ 1 1 1] >-< [ 3 3 1] 0.010100 17 142.0 1.2 46.1 18.4
[ 1 1 1] >-< [ 2 1 0] 0.010710 15 110.4 4.0 30.0 5.9
[ 1 1 1] >-< [ 3 1 1] 0.011070 26 110.4 2.6 19.1 77.9
[ 1 1 1] >-< [ 3 1 1] 0.012770 15 110.4 2.6 19.1 77.9
[ 1 1 1] >-< [ 1 1 0] 0.013260 24 126.2 2.6 40.9 12.1
[ 1 1 1] >-< [ 1 0 0] 0.014880 23 126.2 1.7 8.2 1.1
[ 1 1 1] >-< [ 2 2 1] 0.014880 13 126.2 2.3 10.9 31.1
[ 1 1 1] >-< [ 3 2 1] 0.016000 18 189.3 2.6 26.3 75.4
[ 1 1 1] >-< [ 3 1 2] 0.016000 18 189.3 3.5 19.1 44.6
[ 1 1 1] >-< [ 2 1 0] 0.016150 13 110.4 4.0 30.0 5.9
[ 1 1 1] >-< [ 2 1 1] 0.016420 9 78.9 2.9 30.0 20.8
[ 1 1 1] >-< [ 1 1 0] 0.016510 19 126.2 2.6 40.9 12.1
+----------------------------------------------------------------------------------------+
In this case, the match to the InAs(111) surface with the lowest strain is that with the Al(111) surface, which is in agreement with the experimentally measured structure reported in Ref. [2].
