ALD Precursor Bond Dissociation Energies¶
This application demonstrates an automated workflow to calculate bond dissociation energies (BDE) for atomic layer deposition (ALD) precursor molecules. Given the geometry and atomic symbol of the central atom of an ALD precursor as input, the workflow automatically identifies unique ligand dissociation pathways, generates dissociation reactions, optimizes geometries, and outputs a table of BDE values for all unique ligand types—all with minimal human intervention. This application supports only the first ligand dissociation and is not designed for subsequent dissociation steps.
Important
QuantumATK Version: This application is designed for QuantumATK X-2025.06.
Computational Method: This application uses the Hybrid-DFT level of theory for geometry optimization and energy calculations to achieve accurate BDE predictions.
Automation Level: Fully automated workflow from precursor geometry input to BDE table output.
You can download the required files below:
ald_bond_dissociation_energy_script.py- Automated script for BDE calculationsald_bond_dissociation_energy_workflow.hdf5- Complete automated workflowGenerateFragments.pye- Compile modules for the workflow
Note
The GenerateFragments.pye file must be downloaded and placed in the same directory as the script or workflow file for proper execution.
Key Features
Fully automated workflow: Minimal human intervention from input to output
Automatic ligand identification: Identifies all unique ligand types in the precursor
Automatic reaction generation: Creates dissociation reactions for each unique ligand
Hybrid-DFT calculations: Accurate geometry optimization and energy calculations
Tabular output: Generates an organized table of BDE values for all unique ligand dissociations
Application Goal and Workflow¶
Objective¶
The goal of this application is to calculate bond dissociation energies of ALD precursor molecules in a fully automated manner, requiring minimal human effort.
Input: Geometry of ALD precursor molecule
Output: Table of bond dissociation energies for all unique ligand dissociations
Automated Workflow Steps¶
The workflow executes the following steps automatically:
Read precursor geometry from input file
Identify unique ligands attached to the central metal atom
Generate dissociation reactions for each unique ligand type
Optimize geometries of reactants and products using Hybrid-DFT
Calculate total energies for all optimized structures
Compute BDE values from energy differences
Generate output table with organized BDE results
Scientific Background¶
Bond Dissociation Energy in ALD¶
Bond dissociation energy (BDE) is the enthalpy change required to homolytically break a specific bond in a molecule:
For ALD precursors, BDE provides critical insights into thermal stability and decomposition mechanisms, and guides precursor selection for specific deposition conditions.
Computational Approach¶
The bond dissociation energy is calculated as:
where:
\(E(\text{R-X})\) is the total energy of the intact precursor molecule
\(E(\text{R}^{\bullet})\) is the energy of the first radical fragment
\(E(\text{X}^{\bullet})\) is the energy of the second radical fragment
Automatic Identification of Unique Ligands¶
Critical Automation Step¶
The automatic identification of unique ligands is a crucial step in this workflow. The QuantumATK script analyzes the precursor geometry and automatically determines which ligands are chemically unique, thereby eliminating the need for manual analysis.
Example: Al(CH₃)₂Cl (Dimethylaluminum Chloride)¶
Consider Al(CH₃)₂Cl, a mixed methyl-chloride aluminum precursor:
Ligand Analysis:
Total ligands: 3 (two -CH₃ groups, one -Cl atom)
Unique ligand types: 2 (-CH₃ and -Cl)
Possible First Dissociation Reactions:
Since there are two unique ligand types, there are only two unique dissociation pathways:
Methyl dissociation:
\[\text{Al(CH}_3\text{)}_2\text{Cl} \rightarrow \text{Al(CH}_3\text{)Cl} + \text{CH}_3^{\bullet}\]Chlorine dissociation:
\[\text{Al(CH}_3\text{)}_2\text{Cl} \rightarrow \text{Al(CH}_3\text{)}_2 + \text{Cl}^{\bullet}\]
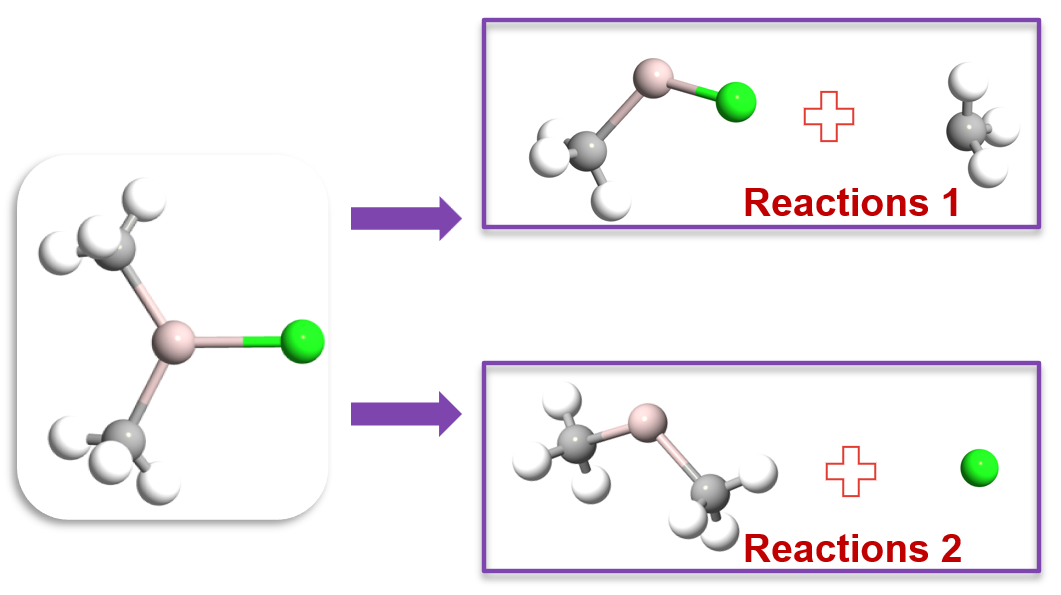
Fig. 152 Illustration of the two unique dissociation pathways for Al(CH₃)₂Cl: (1) methyl group dissociation and (2) chlorine dissociation.¶
Important
Key Principle: The number of unique dissociation pathways equals the number of unique ligand types, regardless of how many copies of each ligand exist.
Two -CH₃ groups are chemically equivalent → analyze only one Al-C bond
One -Cl atom → analyze the Al-Cl bond
Total BDE calculations required: 2 (not 3)
How the Automation Works¶
The automated script performs unique ligand identification by:
Central atom identification: Identify the central metal atom in the precursor based on the atomic symbol provided as user input.
Connectivity analysis: Identifies all ligands bonded to the central metal atom.
Grouping by type: Groups chemically equivalent ligands together based on bond graph analysis and graph isomorphism.
Representative selection: Chooses one representative from each unique group to perform dissociation and generate the dissociation fragments that define the unique reactions.
Automated Output¶
The workflow generates a table of bond dissociation energies:
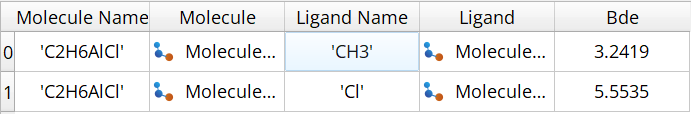
Fig. 153 Example output table showing bond dissociation energies for Al(CH₃)₂Cl calculated using the automated workflow.¶
Summary¶
This automated workflow calculates bond dissociation energies for ALD precursor molecules with minimal human intervention:
Input: Precursor molecule geometry from NanoLab
Process: Fully automated—unique ligand identification, geometry optimization, fragment generation, and BDE calculation using Hybrid-DFT
Output: Formatted table displaying BDE values for all unique ligand dissociations
The automation eliminates manual analysis while providing accurate Hybrid-DFT predictions of bond strengths and decomposition behavior.
