Surface Phase Diagram Construction¶
This application demonstrates how to construct surface phase diagrams for catalytic and reactive surfaces as functions of temperature and gas-phase chemical potentials. Phase diagrams are essential tools for understanding surface stability and reactivity under realistic environmental conditions in heterogeneous catalysis, electrochemistry, and materials science.
Important
QuantumATK Version: This application is designed for QuantumATK X-2025.06.
Module Requirement: This application requires the SurfacePhaseDiagram module for surface phase diagram generation. The module is provided as a compiled Python file (.pye) and must be available in your Python path. You can download it from the attached files.
This application uses thermochemistry data from DFT calculations to construct phase diagrams. You can download the required files below:
SurfacePhaseDiagram.pye- Compiled Python module (Essential - must be in the path)SurfacePhaseDiagram2D.py- Example script for RuO2-H2O-O2 systemSurfacePhaseDiagram2D.hdf5- Workflow for 2D phase diagram generationthermochemistry_database.zip- Thermochemistry database
Key Features
Chemical potential representation: Maps stable surface phases as functions of gas-phase chemical potentials
Temperature-pressure diagrams: Alternative representation showing phase stability vs. temperature and pressure
Multi-species support: Handles binary gas environments (e.g., H2O and O2, CO and O2)
Multiple phases: Determines most stable configuration among clean and various adsorbate-covered surfaces
Flux calculations: Provides molecular flux information for experimental comparison
Customizable plots: Generates high-quality phase diagrams
Scientific Background¶
Surface Phase Diagrams in Catalysis¶
Surface phase diagrams provide crucial insights into:
Catalyst active phases: Which surface termination is stable under reaction conditions
Reaction mechanisms: How surface composition changes affect catalytic activity
Poisoning and deactivation: Understanding conditions where unwanted species dominate
Operando conditions: Relating in-situ observations to surface structures
Theoretical Framework¶
The phase diagram is constructed by calculating the surface free energy (γ) as a function of chemical potentials (μ) for each gas-phase species:
where:
A is the surface area per unit cell
Gsurf is the Gibbs free energy of the surface configuration
Gbulk is the Gibbs free energy per formula unit of bulk material
nbulk is the number of bulk formula units in the surface slab
nspecies are the numbers of adsorbed molecules
μspecies are the chemical potentials of gas-phase species
The chemical potential relates to pressure through:
At each point in (μ₁, μ₂) space, the phase with the lowest surface free energy is thermodynamically stable.
Example System: RuO2 in H2O/O2 Environment¶
The example system demonstrates phase diagram construction for RuO2(110), a model oxide catalyst, in contact with water vapor and oxygen gas. This system is relevant for:
Surface is in equilibrium with humid environments
Multiple gaseous species influence surface stability
Surface Configurations¶
The phase diagram includes multiple competing surface terminations:
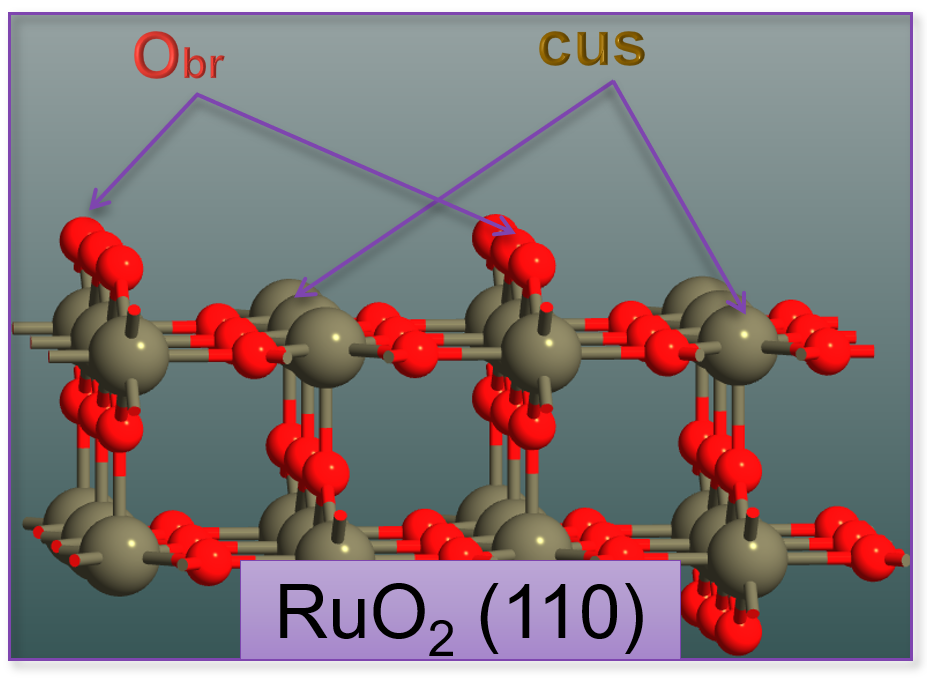
Fig. 150 RuO2(110) surface structure showing key adsorption sites. The surface features coordinatively unsaturated Ru atoms (cus sites) and bridge oxygen atoms (br sites) that serve as active sites for molecular and dissociative water adsorption.¶
- 1. Clean Surface (Obr/-)
The bare RuO2(110) surface with coordinatively unsaturated Ru and bridge oxygen sites
- 2. Molecular Water Adsorption (H2O/cus)
Water molecules physisorbed on coordinatively unsaturated (cus) Ru sites
- 3. Dissociated Water (2OH/cus+br)
Water dissociated into OH groups on both cus and bridge (br) sites
- 4. Hydroxylated + Water (OH + H2O)
Mixed phase with both OH groups and molecular water
- 5. Oxygen Terminations (O/cus, 2O)
Additional oxygen atoms adsorbed on the surface from O2 dissociation
Workflow Overview¶
The complete workflow consists of six main steps:
STEP 1: Generate Surface Configurations
↓
STEP 2: Calculate DFT Energies & Thermochemistry
↓
STEP 3: Store Thermochemistry in Database
↓
STEP 4: Define System Parameters
↓
STEP 5: Configure Phase Diagram Script
↓
STEP 6: Generate Phase Diagram
STEP 1: Generate Surface Configurations¶
Use QuantumATK’s Builder tools to create all relevant surface structures:
Create clean surface:
Use Surface (Cleave) tool to generate the desired surface orientation
Optimize slab thickness (typically 3-5 atomic layers plus vacuum)
Add appropriate vacuum spacing (>10 Å)
Generate adsorbate configurations:
Open the Adsorption Tool (Builder → Surface Tools → Add Adsorbates)
Add water molecules at different sites (atop, bridge, hollow)
Create dissociated configurations (OH, O species)
Consider different coverages (1/2, 1/4, full monolayer)
Save all configurations as separate structure files for subsequent calculations
Tip
For systematic exploration, generate a comprehensive set of possible adsorption configurations. The phase diagram will automatically identify which are stable under different conditions.
STEP 2: Calculate Thermochemistry Properties¶
Compute thermochemistry data for all species using appropriate workflow templates:
For Gas-Phase Species (H2O, O2):
# Use IdealGasThermoChemistry workflow
# This calculates:
# - Optimized geometry
# - Electronic energy
# - Vibrational frequencies
# - Rotational and translational contributions
# - Temperature-dependent free energy
For Bulk and Surface Configurations:
# Use CrystalThermoChemistry workflow
# This calculates:
# - Optimized crystal structure
# - Electronic energy
# - Phonon density of states (optional for T > 0)
# - Temperature-dependent free energy
Note
Computational settings recommendations:
Use consistent DFT functional for all calculations (e.g., PBE)
Converge k-point sampling and basis set cutoffs
For surfaces: Use sufficient slab thickness and vacuum
Include spin polarization where relevant (O2, transition metal oxides)
STEP 3: Upload Thermochemistry to Database¶
Store computed thermochemistry data in QuantumATK’s thermochemistry database:
Access the database:
Open Thermochemistry Analyzer in NanoLab
Click Database button to open database window
Default location:
C:\Users\username\.quantumatk\databases\thermochemistry\
Create entries:
Click Add Entry for each calculated configuration
Provide unique, descriptive names (e.g., “RuO2_bulkDFT”, “RuO2_OH_cusDFT”)
Upload the corresponding thermochemistry object from your calculation files
Organize entries:
Use systematic naming conventions
Document stoichiometry and configuration details
Keep track of all entry names for the script configuration
Alternatively:
Open the database folder (default location:
C:\Users\username\.quantumatk\databases\thermochemistry\)Copy the thermochemistry objects directly into the folder with unique names
Note: No multipliers will be set when using this method
STEP 4: Define System Parameters¶
Set the physical parameters for your phase diagram at the top of the script:
# Temperature and pressure reference
temperature = 100.0 * Kelvin # Operating temperature
p0 = 1.013 * bar # Reference pressure (1 atm)
# Chemical potential ranges for species 1 (e.g., H₂O)
mu_species1_min = -2.5 * eV
mu_species1_max = 0.5 * eV
# Chemical potential ranges for species 2 (e.g., O₂)
mu_species2_min = -2.5 * eV
mu_species2_max = 0.0 * eV
# Molecular masses for flux calculations
mass_species1 = 18.015 * atomicMassUnit # H₂O
mass_species2 = 32.0 * atomicMassUnit # O₂
Tip
Choosing chemical potential ranges:
Typical range: -3 to 0 eV relative to gas-phase reference
Lower μ: low pressure/activity conditions
Higher μ: high pressure/activity conditions
Adjust based on relevant experimental conditions
STEP 5: Configure Script Parameters¶
Edit the script to match your system by updating three main configuration dictionaries:
A. SYSTEM_ENTRIES Dictionary
Define all database entry names and stoichiometry:
SYSTEM_ENTRIES = {
# Bulk reference
'bulk_config': 'RuO2_bulkDFT', # Database entry name
'bulk_stoichiometry': 2.0, # Formula units in bulk
# Gas phase references
'gas_species2': 'O2DFT', # O₂ gas thermochemistry
'gas_species1': 'H2ODFT', # H₂O gas thermochemistry
# Clean surface
'clean_surface': 'RuO2_O_brDFT', # Clean surface entry
'surface_stoichiometry': 8.0, # Bulk units in slab
'clean_surface_label': 'Obr/-', # Plot label
# Adsorbate configurations
'surf_ad_configs': [
{
'name': 'RuO2_OH2_cusDFT', # Database entry
'label': 'H\ :sub:`2`\ O/cus', # Plot label
'color': 'skyblue', # Phase color
'n_species1': 1.0, # Number of H₂O
'n_species2': 0.0, # Additional O atoms
},
{
'name': 'RuO2_2OH_cus_brDFT',
'label': '2OH/cus+br',
'color': 'lightgreen',
'n_species1': 1.0, # From dissociated H₂O
'n_species2': 0.0,
},
# Add more configurations...
],
}
B. PHASE_DIAGRAM_CONFIG Dictionary
Set the phase diagram grid and display parameters:
PHASE_DIAGRAM_CONFIG = {
# Chemical potential grid
'del_mu_species2_range': (
mu_species2_min.inUnitsOf(eV),
mu_species2_max.inUnitsOf(eV),
0.01, # Step size in eV
),
'del_mu_species1_range': (
mu_species1_min.inUnitsOf(eV),
mu_species1_max.inUnitsOf(eV),
0.01,
),
# Pressure axis tick positions
'pressure_ticks_species2': (
mu_species2_min.inUnitsOf(eV),
mu_species2_max.inUnitsOf(eV),
0.5, # Tick spacing
),
'pressure_ticks_species1': (
mu_species1_min.inUnitsOf(eV),
mu_species1_max.inUnitsOf(eV),
0.5,
),
'reference_pressure': p0,
'title': 'RuO\ :sub:`2`\ (110) Surface Phase Diagram',
'output_filename': 'surface_phase_diagram.png',
}
C. PLOT_CONFIG Dictionary
Customize plot appearance:
PLOT_CONFIG = {
'figure_size': (18, 16), # Width, height in inches
'dpi': 300, # Resolution
'title_fontsize': 16,
'label_fontsize': 16,
'tick_fontsize': 14,
'phase_label_fontsize': 12,
'clean_surface_color': 'violet', # Color for clean surface
}
STEP 6: Run the Phase Diagram Script¶
Execute the script to generate the phase diagram:
# Import the module
import SurfacePhaseDiagram as surface_phase_diagram
# Generate the plot
fig, stable_phase_index, thermo_data = surface_phase_diagram.generate_surface_phase_diagram(
SYSTEM_ENTRIES,
PHASE_DIAGRAM_CONFIG,
PLOT_CONFIG,
MOLECULAR_MASSES,
temperature,
p0,
)
The script will:
Load all thermochemistry data from the database
Create a 2D grid in (μ₁, μ₂) space
Calculate surface free energy for each phase at each grid point
Determine the most stable phase at each point
Generate a color-coded phase diagram with multiple axes
Save the plot as a high-resolution PNG file
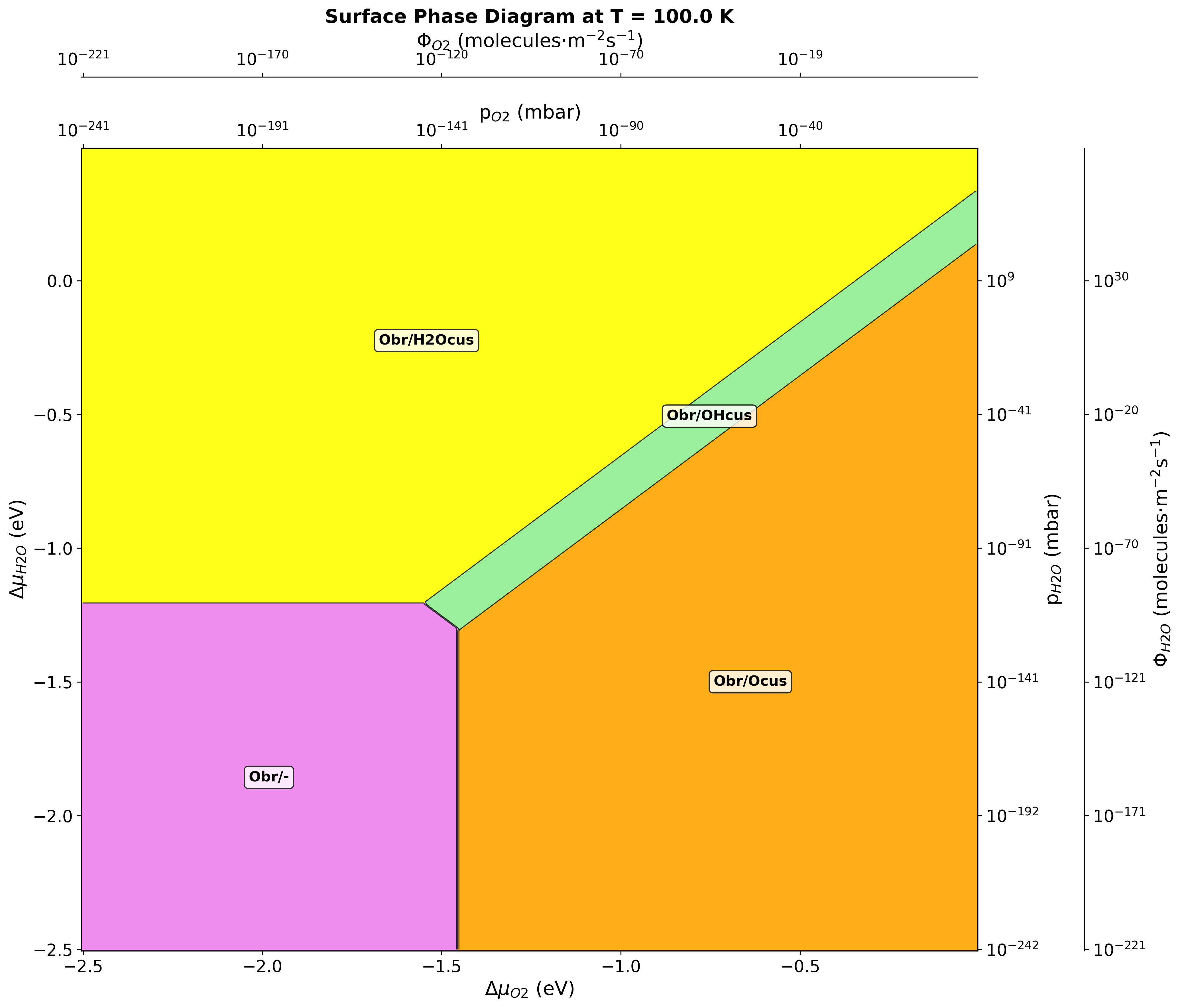
Fig. 151 Surface phase diagram for RuO2(110) in H2O-O2 environment at 100 K. Different colors represent stable phases: clean surface (violet), molecular water adsorption (yellow), dissociated water (green), and oxygen-rich terminations (red/orange).¶
Temperature-Pressure Phase Diagrams¶
Alternative Representation¶
Instead of chemical potentials, phase diagrams can be plotted as temperature vs. pressure:
# Generate T-P diagram instead of μ₁-μ₂ diagram
fig, stable_phase_index, thermo_data = surface_phase_diagram.generate_temperature_pressure_diagram(
SYSTEM_ENTRIES,
TEMP_PRESSURE_CONFIG,
PLOT_CONFIG,
)
Configuration for T-P diagrams:
TEMP_PRESSURE_CONFIG = {
'temperature_range': (100, 1000, 10), # Min, max, step in K
'pressure_range': (1e-10, 1e5, 100), # Min, max, steps in bar
'fixed_gas_species': 'gas_species1', # Which gas to vary
'fixed_gas_pressure': 1e-3 * bar, # Pressure of fixed gas
'output_filename': 'phase_diagram_TP.png',
}
This representation is more intuitive for experimental comparison and directly shows operating windows for catalytic processes.
Advanced Usage¶
Multiple Temperature Comparison¶
Generate phase diagrams at different temperatures to understand temperature effects:
temperatures = [100, 300, 500, 800] * Kelvin
for T in temperatures:
fig, _, _ = surface_phase_diagram.generate_surface_phase_diagram(
SYSTEM_ENTRIES,
PHASE_DIAGRAM_CONFIG,
PLOT_CONFIG,
MOLECULAR_MASSES,
temperature=T,
p0=p0,
)
# Save with temperature-specific filename
Running the Script¶
The complete script can be executed in several ways:
From NanoLab:
Open the script file in the Scripteditor
Click Send To → Job Manager
Or select the file and use Jobs tool to execute
From Command Line:
atkpython SurfacePhaseDiagram2D.py
Note
Running the Script Locally
This script requires access to the thermochemistry database containing all computed thermochemistry objects for the bulk, gas-phase species, clean surface, and adsorbate configurations. The script must be executed locally on a system where:
The thermochemistry database is available and properly configured
All required thermochemistry entries have been calculated and uploaded to the database
The SurfacePhaseDiagram.pye module is in the Python path
Output Files¶
The script generates:
surface_phase_diagram.png: The complete phase diagram with all axes
Console output: Progress information and verification messages
The phase diagram image includes:
Color-coded phase regions
Chemical potential axes (eV)
Pressure scales (mbar)
Molecular flux scales (molecules·m⁻²·s⁻¹)
Phase labels
Temperature and title information
Further Reading¶
Theoretical Background:
Reuter, K., & Scheffler, M. (2001). “Composition, structure, and stability of RuO2(110) as a function of oxygen pressure.” Physical Review B, 65(3), 035406.
Reuter, K., & Scheffler, M. (2003). “Composition and structure of the RuO2(110) surface in an O2 and CO environment: Implications for the catalytic formation of CO2.” Physical Review B, 68(4), 045407.
Knapp, M., Crihan, D., Seitsonen, A. P., Lundgren, E., Resta, A., Andersen, J. N., & Over, H. (2006). “Effect of a humid environment on the surface structure of RuO2(110).” Journal of Physical Chemistry B, 110(30), 14007-14010.
Summary¶
This application enables construction of surface phase diagrams from first-principles thermochemistry calculations. Key capabilities include:
✓ Map stable surface phases as functions of gas-phase chemical potentials or T-P conditions
✓ Support for multi-species environments and complex surface configurations
✓ Flexible framework adaptable to different material systems and conditions
Phase diagrams provide essential insights for understanding and predicting surface behavior under realistic environmental conditions, bridging the gap between computational predictions and experimental observations in catalysis, electrochemistry, and materials science.
